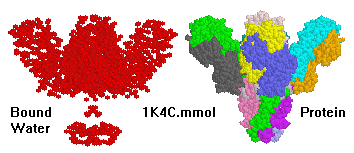
Macromolecules
for Protein Explorer
Suggestions to Eric Martz.
All images copyright © 2002 by Eric Martz.
Click on any image for more information.

| Macromolecules for Protein Explorer Suggestions to Eric Martz. All images copyright © 2002 by Eric Martz. Click on any image for more information. |
|
Four ways to use the PDB ID codes below:
To find the primary literature citation for a molecule, the full name and species from which it came, etc., after displaying the molecule in PE, go to PE's Features of the Molecule. Years in parentheses after links to molecules indicate the years of publication. In some cases a range of years is given: the early year is when the molecule was first solved (if I happen to know that) at 3.5 Å resolution or better; the later year is when the structure chosen for the Atlas was published. |
In each category below, PDB files have been divided into those that are relatively straightforward, those that are more challenging, and sometimes enormous. "Straightforward" cases have been selected to avoid complications (such as being NMR ensembles, lacking sidechains, having a nonstandard file format, having many alternative sidechain conformations, etc.). Lesson Plans: It is best if your students first have one to two hours to do the 1-Hour Tour to become familiar with Protein Explorer (PE). Then you can assign molecules from the list below to each group of students (or let them pick), and hand out Discovery in Protein Explorer, a generic set of questions, some open-ended. Optionally, you can provide hints suggesting how to use PE to answer the questions. If you use PE in your class, please do the Student Assessment of Learning Gains. Acknowledgements. Some cases in the Atlas came from Tim Herman and Michael Patrick's 2001 SEPA Course. Thanks to PDB Files for Teaching Biochemistry by Don Harden and Dabney Dixon of Georgia State University, and Molecule of the Month by David Goodsell for some of the cases below. For suggestions that have been incorporated below, thanks also to Ilan Samish, David Margulies, and Bruce Southey. Thanks to the EBI Probable Quaternary Structures server by Kim Henrick and Janet Thornton which has been invaluable in many cases below. |
| Enzymes |

 CHALLENGING
CHALLENGING
| Signal Cascade Proteins (Cytoplasmic) |
| Soluble Proteins (Not Enzymes) |

See (under other categories)
CHALLENGING

| Toxins |
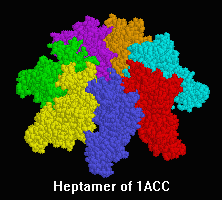 Anthrax Toxins
Anthrax Toxins
| Structural Proteins |

| Calcium-Binding Proteins |
See (under other categories)
CHALLENGING
| Lipid Bilayers |
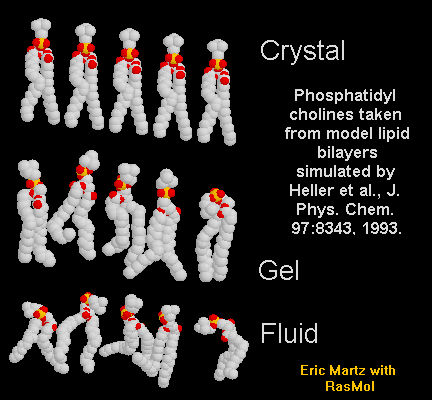 (Yes, we know they're not really macromolecules.)
(Yes, we know they're not really macromolecules.)
| Integral Membrane Proteins |



CHALLENGING

| Myristoylated Proteins |
 |
See also (under other categories)
| DNA and RNA |
Genes were shown to reside in DNA in 1944 (Avery et al.) and this became widely accepted after the 1952 experiments of Hershey and Chase. The double helical structure of the DNA was predicted by James Watson and Francis Crick in 1953 (Nobel Prize, 1962). Their prediction was based in part upon X-ray diffraction studies by Rosalind Franklin, to whom Watson and Maurice Wilkins gave inadequate credit (see Rosalind Franklin: Dark Lady of DNA by Brenda Maddox, HarperCollins, 2002). The predicted B-form double helix was not confirmed with atomic-resolution crystal structures until 1973, first by using dinucleotides of RNA (Rosenberg et al.). The first crystal structure containing more than a full turn of the double helix was not solved until 1980 (Wing et al. 1BNA, 1981, 12 base pairs). The lag of more than a quarter century between prediction and empirical confirmation involved development of X-ray crystallography for macromolecules, and the need to produce a short, defined sequence of DNA for crystallization. This brief account is based upon a review by Berman, Gelbin, and Westbrook (Prog. Biophys. molec. Biol. 66:255, 1996), where the references will be found.



 Transfer RNA (Phe),
4TNA
(1974-1978).
(1EHZ
[2000] is a more challenging tRNA.)
Transfer RNA (Phe),
4TNA
(1974-1978).
(1EHZ
[2000] is a more challenging tRNA.)
CHALLENGING
| Proteins Complexed to Nucleic Acids (Transcription Factors, Polymerases, etc.) |

CHALLENGING
| Virus Capsids |
ENORMOUS

| Magnificent Molecular Machines |
CHALLENGING

SEE ALSO (under other categories)
| Immune System & Defense Molecules (Antibody, etc.) |

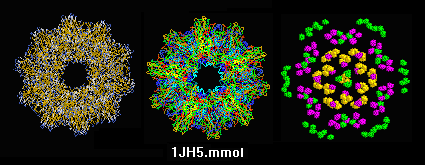 sTALL-1 cytokine of the TNF superfamily,
human, soluble fragment expressed in E. coli,
1jh5_cao.mmol
(2001; alpha carbons only).
sTALL-1 cytokine of the TNF superfamily,
human, soluble fragment expressed in E. coli,
1jh5_cao.mmol
(2001; alpha carbons only).

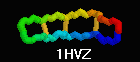 Defensin (Rhesus theta defensin one, RTD-1), an antibacterial
cyclic 18-amino acid peptide,
1HVZ (20-model NMR strcture, 2001).
Each half is coded by a different gene. Cyclization is thought to
increase resistance to exoproteases
(Trabi & Craik, TiBS 27:132).
Defensin (Rhesus theta defensin one, RTD-1), an antibacterial
cyclic 18-amino acid peptide,
1HVZ (20-model NMR strcture, 2001).
Each half is coded by a different gene. Cyclization is thought to
increase resistance to exoproteases
(Trabi & Craik, TiBS 27:132).
CHALLENGING
| Carbohydrates |
| Unusual Tertiary and Quaternary Structures |

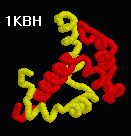

| Animated Morphs of Conformational Changes |
| Evolutionary Conservation |
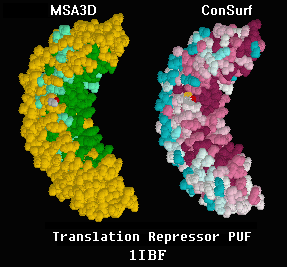
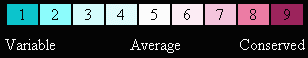

| Protein Crystals |

| History: Earliest Crystallographic Structures |


| Other Browsable Lists of Molecules |
Suggestions to Eric Martz.
* PDB file sizes marked "*" are given for
gzipped files, as they will be transferred
from the Protein Data Bank (or other servers)
to Chime when the above links are clicked.
If you save the plain text PDB file to disk from Chime/PE, it will
be about 4-fold larger.
Alpha-carbons only:
Some very large structures are supplied as alpha-carbons only. This enables
the backbone to be viewed, but not secondary structure (and therefore
also not
schematic "cartoon"
rendering). To see secondary structure, you'll have to get the complete
structure (usually available from EBI's
Probable Quaternary Structures), but it is typically multiple megabytes
in size.