Visualizing Regions of Conservation
in 3D Protein Structures
|
|
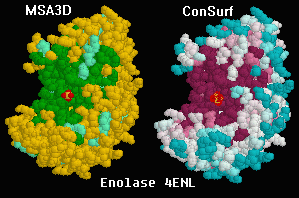
Dark regions show conservation in
the catalytic site of enolase.
|
Regions of evolutionary conservation or variability
(lower or higher than average mutation rates, respectively)
can be visualized on a 3D protein structure
by applying a color scheme based upon a multiple sequence
alignment. Such regions generally signal clusters of residues
of crucial functional importance.
In December, 2001, Glaser, Ben-Tal, Pupko and
Martz released the
ConSurf Server, where you will find a
Gallery of
exemplary results. ConSurf automatically generates a
multiple protein sequence alignment (or accepts one you have made),
then automatically generates a phylogenetic tree,
and applies colors representing the resulting grades of conservation
for each residue to the 3D protein structure. The results
are displayed in Protein Explorer.
Earlier, beginning in 2000, Protein Explorer
offered MSA3D, which accepts a user-provided multiple protein sequence
alignment, and uses it to color the 3D protein structure. MSA3D
simply divides residues into 3 categories: identical, similar, and different.
ConSurf is much easier to use than MSA3D, because it is completely automatic
and provides everything needed at a single site.
More importantly, the algorithms employed by ConSurf are much more sophisticated.
For nearly all purposes, ConSurf is superior to MSA3D. Therefore we
strongly recommend that you use
ConSurf to visualize regions of conservation in 3D protein structures.
The documentation below, describing MSA3D, is now mostly of historical
interest. Protein Explorer's MSA3D routines remain available in case they
are useful for specialized purposes.
Protein Explorer's Multiple Sequence
Alignment in 3D (MSA3D): Enolase
A multiple protein sequence alignment was prepared for enolase.
Protein Explorer's MSA3D feature then assigned colors
to represent identity, similarity, or difference
of the aligned amino acids, shown in Protein Explorer's
alignment listing.
The alignment includes eubacteria, archebacteria,
and eukaryotes (Drosophila, yeast, and human). Despite
this enormous span of evolutionary time, all of the residues
in the catalytic site pocket are identical. (The catalytic site
is marked by the red sulfate ion that happens
to be bound there in this structure, 4enl.)
If you have Chime installed, you can
see this molecule rotate in an interactive window.
If not,
downloading Chime and installing it takes only
a few minutes.
The links and buttons in the snapshot below don't work since this
is just a snapshot.
Thanks to Paul Stothard for major portions
of the MSA3D code, and to Garry Duncan for providing this alignment.
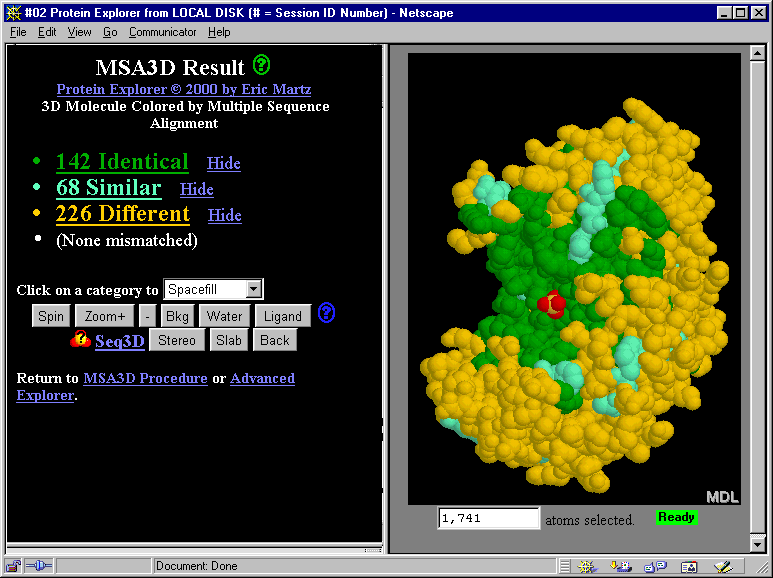
How do you find MSA3D after you start Protein Explorer?
If you start Protein Explorer from the special link below, you'll
see a link to MSA3D immediately. Most other links to Protein Explorer
will first give you a FirstView description.
From FirstView, click "Explore More", and there you'll see
a link to "Advanced Explorer", where you'll see MSA3D on the menu.
The enolase alignment is a built-in demonstration in MSA3D.
This special
Enolase-MSA3D link
starts Protein Explorer, loads enolase,
then automatically changes from FirstView directly
to Advanced Explorer, where the MSA3D
link will be evident.
A
detailed tutorial is provided on how to construct an alignment
and use it to color a molecule of interest with MSA3D.


